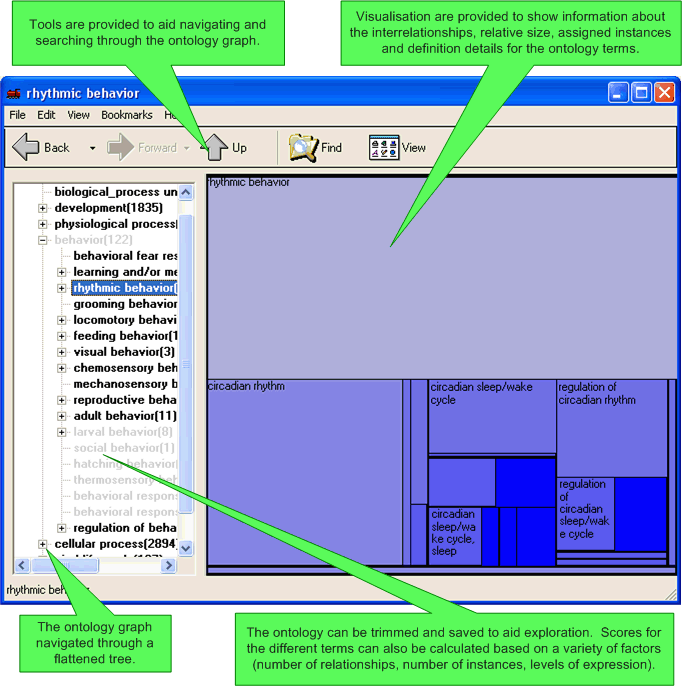
SeqExplorer is an add-on to SeqExpress which allows for the visualisation, scoring and browsing of complex ontologies, specifically the Gene Ontology. It provides a number of visualisation tools for the analysis of interrelationships within the ontology, examination of scores associated with concepts (e.g. number of assigned instances), viewing and editing of the specific concept details and concept similarity comparison (for the identification of possible errors). Data importing, scoring and searching tools are also provided.
When SeqExplorer is first loaded it will automatically show a 'slim' version of the biological process ontology with human gene annotations, other ontologies and annotations can then be loaded.

SeqExplorer's menu system provides the following functionality:
File: for the saving and loading of ontology information.
New Browser: this will bring up a new SeqExplorer window.
Open...: this will open a saved SeqExplorer file.
Save/Save As: saves the current information to a data file.
Import...: Used to import either an ontology in OBO format, or a tab separated file of instances (proteins) that have been classified with specific ontology terms.
Export: Used to either export the ontology to OBO format, or to export a tab separated file of instances (proteins) that have ben classified with specific ontology terms. The export is done from the current selected ontology term downwards (presuming the ontology is a directed graph) following all active relationships, it is thus possible to save sub ontologies and prune the graph.
Edit: for editing both the ontology terms and defining scores.
Rename: Renames the ontology term.
Scores: choose the scoring method that is to be used. Three scoring methods are available: Relationship (counts the number of child concepts in the direct graph for each concept), Instance( counts the number of classified instances that have been marked up with each concept), Expression (uses the values from a gene expression experiment - this option requires a full SeqExpress installation to be present).
Relationship Scores. Configures how the scoring using the number of relationships is performed. Can choose whether or not to follow different relationships types (e.g. is_a, part_of, has_a) to calculate the number of 'child' concepts each concept has.
Instance Scores. Configures how the scoring using the number of instances is performed. Can choose whither or not to count a specific instance depending on its evidence code (e.g. IEA).
Expression Scores Configures how the scoring using data from a (set of) gene expression experiments is performed. Either the maximum value the,mean value or the minimum value of the expression profiles associated with each concept are used to calculate the score.
Find. Enables the searching through the name of each concept and any textual descriptions associated with them.
View: A number of different visualisations are supported in SeqExplorer
TreeMap: Visualises the scores and relative 'sizes' of each of the concepts in the ontology.
Graph:. Shows the interrelationships between the ontology terms.
Matrix: Shows the similarity between instances that have been annotated with different ontology terms.
List: Shows the details of a specific concept.
BookMarks. Creating and editing bookmarks to concepts of interest.
Add. Will add the currently associated concept (and active relationship) to the bookmark list.
Organise. Use to alter the order, rename or delete the bookmarks.